How the microbiome is the next chapter in precision medicine
Discover how genome sequencing and metagenomic analysis are helping scientists advance precision medicine

Principal Strategic Opportunities Manager

The Human Microbiome; A new frontier in drug discovery.
What is the microbiome, and why is it important?
The human body is home to a huge and diverse community of microorganisms, consisting of bacteria, fungi, protozoa and viruses. Collectively these organisms are referred to as the microbiota, and the collection of genomes from these microorganisms is referred to as the microbiome, however the two terms are often used interchangeably.
It is estimated that over 10,000 species of microorganisms occupy the human microbiota, and that there may be more than eight million unique genes associated with the various microbiomes in the human body. The number of microorganisms present in the human microbiota is thought to be slightly higher than that of human cells in the body, and some have classified the microbiome as an organ in its own right1.
Microorganisms inhabit many parts of the human body, including the gastrointestinal tract, skin, saliva, oral mucosa and conjunctiva, with the vast majority residing in the colon. Each of these microbiomes is distinct depending on the location in the body, and the relationship is generally mutually beneficial.
The microbiome plays a critical role in human health, supporting processes such as:
- Maintaining intestinal integrity and supporting barrier function
- Breaking down food
- Providing essential vitamins, nutrients and beneficial short-chain fatty acids such as butyrate
- Regulating the immune system, for example, modulation of lymphoid structure development and T cell differentiation 2
- Protecting against infection through secretion of bacteriocins and antagonising pathogens
- Contributing to mental health
- Regulating metabolism
There has been an explosion in our understanding of the human microbiome in recent years as advances in genome sequencing technologies and metagenomic analysis have enabled scientists to study these communities of microorganisms.
Tremendous variation can be observed in terms of the species that make up a person’s microbiome, and even within an individual, these microbial communities can shift depending on factors such as diet, medication, age, stress levels and other environmental factors.
The microbiome has become a hot topic for research recently, as when the balance of the microbiota is disturbed the microbiome can contribute to a range of diseases and conditions including cancer, diabetes, inflammatory bowel disease, autism, anxiety and obesity.
Analysis of the microbiome can be performed using DNA sequencing techniques, including 16S rRNA sequencing and shotgun metagenomics. The sequence of the 16S rRNA gene consists of highly conserved sequences which alternate with nine windows of hypervariable regions, enabling universal PCR primers to be used to target the conserved regions to sequence many species at a time, which can then be distinguished based on the sequence of the variable regions.
16S rRNA gene sequencing provides a robust tool for identifying and classifying the bacterial species within a sample, however, is limited in that it does not give information on the overall ecology of the microbiome and the interactions that are occurring. Shotgun metagenomics can overcome these limitations to some extent, providing a higher taxonomic resolution and the possibility to extract the functional gene content of each genome and aiding understanding of how the microbiome is functioning3.
Microbiome research is changing our perception of human biology and it is now becoming apparent that we must consider the microbes with which we have coevolved as a critical factor in health and disease.
The gut microbiome and gut-brain axis
To date a major focus of human microbiome research has been studying the bacteria in the gut. The gut microbiota represent the largest and most diverse community in the human microbiome. The gut microbiota is dominated by strictly anaerobic organisms, with members of the Firmicutes and Bacteroidetesphyla as the most dominant species followed by Proteobacteria and Actinobacteria.4.
The adult gut microbiota is resilient against minor perturbations, however more significant influences such antibiotic treatment or disease onset can lead to dramatic changes in the microbial community composition. An individual’s genetics, diet and environment also play a major role in shaping the gut microbiota5.
The gut microbiome plays an important role in metabolism and regulating the immune response, but its impact can also be observed on the brain, where microbial metabolites influence mood and behaviour via pathways that connect the gut to the central nervous system, including direct activation of the Vagus nerve from the enteric nervous system, production of metabolites that pass through the intestinal barrier into the circulatory system, and modulation of the immune system through metabolites and microbial associated molecular patterns6.
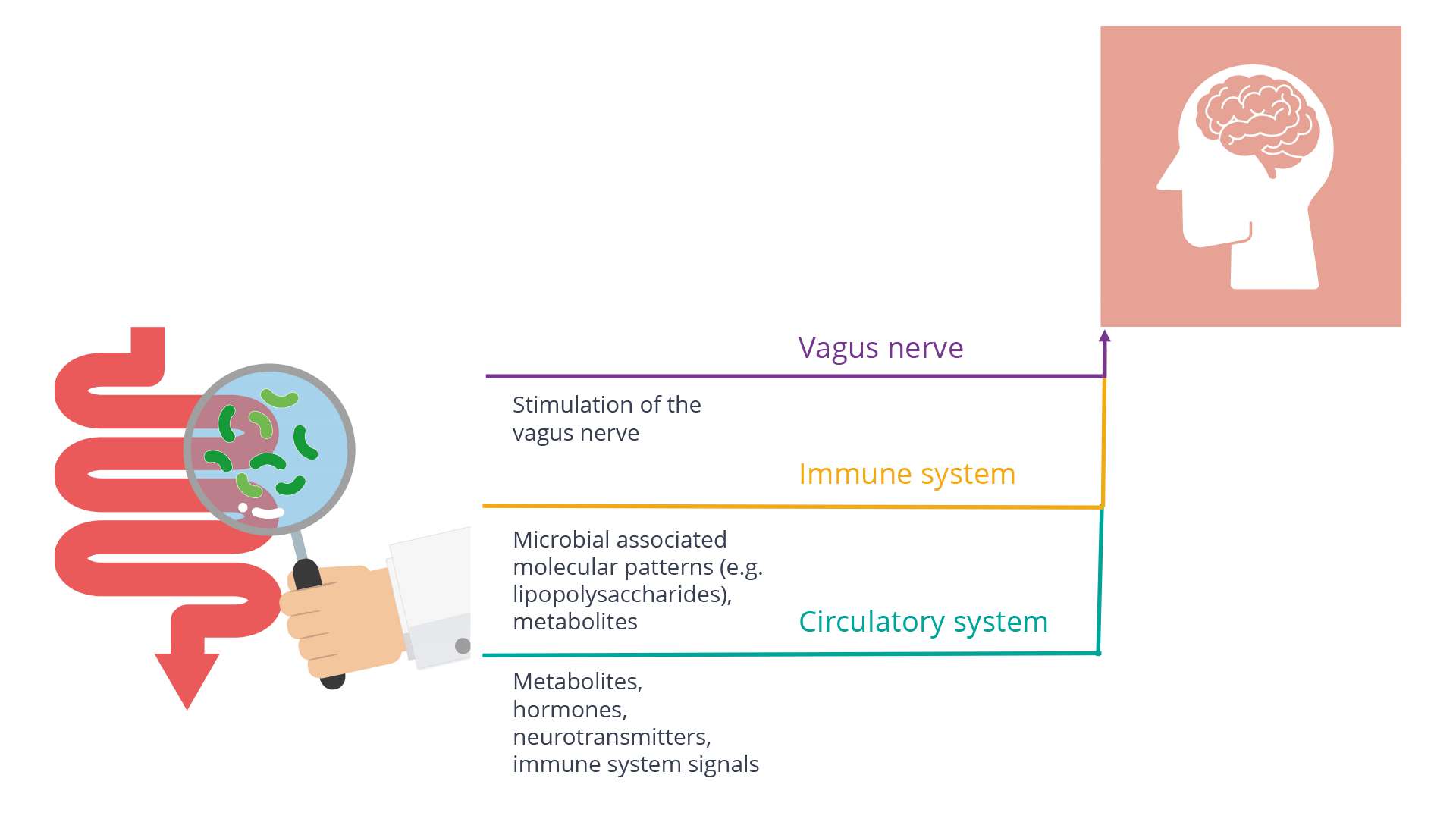
There is increasing recognition that psychiatric and neurological illnesses frequently co-occur with gastrointestinal pathology, and studies indicate that the composition of the host microbiome affects neurological function and leads to effects including depression and anxiety, and treatment with certain species of Lactobacillusor Bifidobacterium probiotics can reduce anxious and depressive behaviour 7, 8, 9.
As we begin to understand how the microbiome influences the brain and behaviour this opens up possibilities for rationally designed therapeutics to treat mental health illnesses that are currently treated by pharmacological alteration of neurotransmitters, for example selective serotonin reuptake inhibitors that are commonly used to treat depression.

Microbiome therapeutics
Therapeutic manipulation of the microbiome is a rapidly advancing field and data suggests that treatments capable of reversing a damaged, dysbiotic microbiome are effective in managing certain human diseases. Therapeutic approaches used to modify the microbiome may include:
- Faecal microbiota transplants (FMT) to rebalance the gut microbiome, a technique already proven to be extremely successful in tackling Clostridium difficile infections and one of the most direct approaches to manipulate the microbiome
- Probiotic supplements, e.g. Lactobacillus and Bifidobacterium species to recolonise the patient with beneficial microbial species. Probiotic treatment could potentially include rationally designed therapeutics containing a personalised consortium of microorganisms dependent on the patient’s microbiome.
- Subtractive microbiome therapeutics to eliminate deleterious members of the microbiome, for example using chemical, peptides, bacteriophages or bacteriocins
- Prebiotic supplements to modulate the composition or activity of the microbiome. Prebiotics aim to increase the levels of beneficial bacteria and may include non-digestible fibre components (for example inulin or oligosaccharides) to stimulate the growth of Bifidobacterium and other beneficial microorganisms. Prebiotics are found naturally in human milk and are now added to many commercial formulas of infant milk.
- Synbiotics, a combination of prebiotics and probiotics
- Microbial vaccines, for example, Merck have partnered with the microbiome company 4D Pharma to develop vaccines based on delivering live bacteria to the gut
- Next-generation therapeutics will likely include recombinant probiotics with engineered characteristics and controllable gene circuits, as this offers an opportunity to improve existing microbial communities
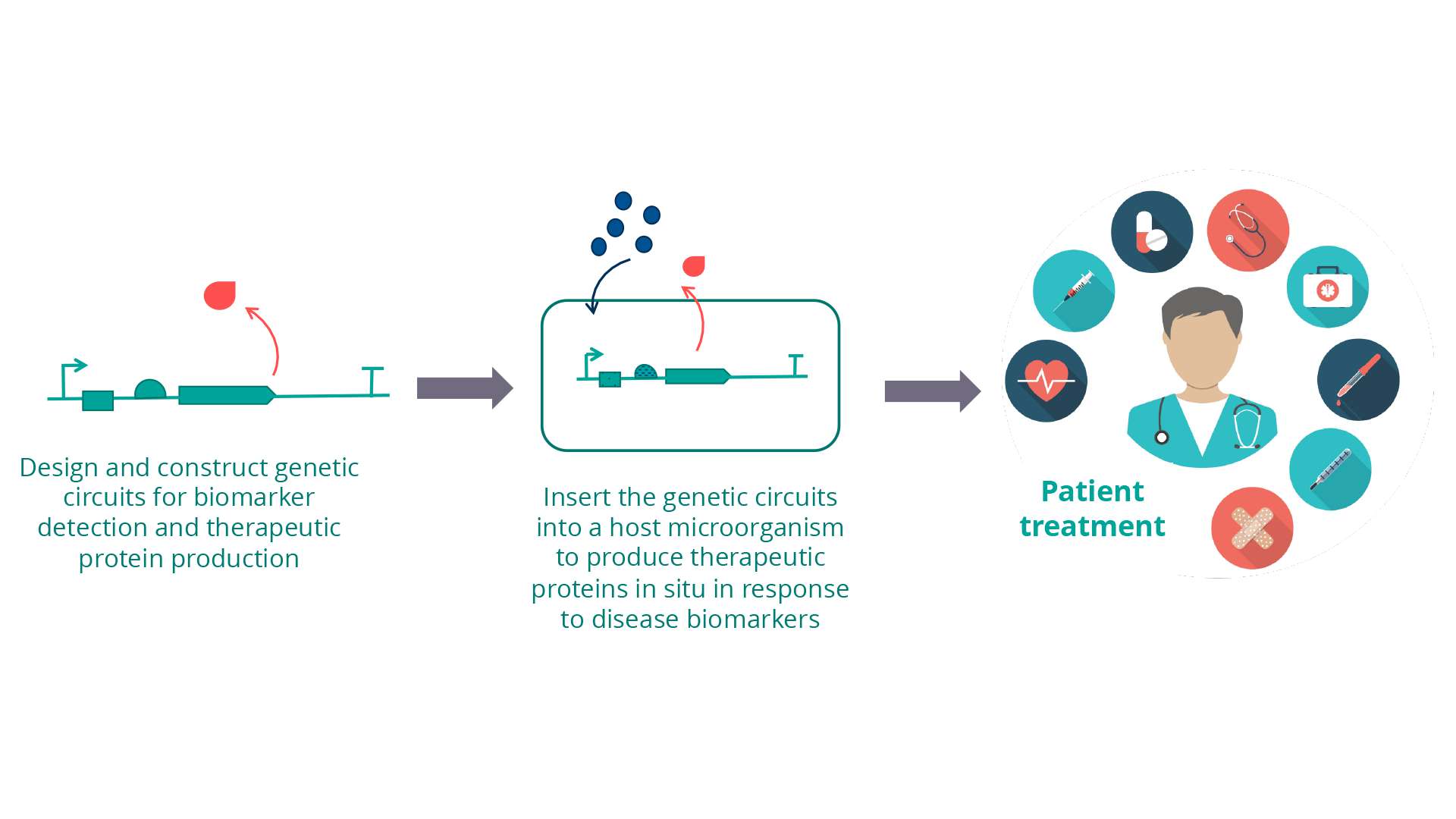
There is also potential in the future to use the microbiome as a drug delivery system, for example through the use of engineered probiotics containing genetic circuits to enable the production of therapeutic proteins upon detection of disease biomarkers, enabling on-demand drug release10.
As we move forward with efforts to manipulate the microbiome to enhance health and therapeutic responses there is still a lot to learn around the mechanisms by which the microbiome exerts its effect.
Understanding the factors that govern the composition of the gut microbiota will be of critical importance in designing successful microbiome therapeutics, and developing microbiome therapies that are robust enough to withstand the diversity and an individual’s microbiome will be a major challenge 10.
One of the biggest challenges in microbiome research is determining whether a change in the microbiota is responsible for a specific condition, or if it is a side effect of having the condition. The complex nature of the microbiome and the fact that every person has a distinct one makes it challenging to determine cause-effect relationships.
The microbiome as the next chapter in precision medicine
The paradigm of precision medicine promises to help improve outcomes and reduce adverse effects of treatment, improve preventative care and reduce healthcare costs by tailoring treatment to an individual.
The sequencing of the human genome has led to advances in our understanding of the genomic basis of disease, and significant advances towards treating patients at a more individual level based on their genetic information and known biomarkers for disease.
The field of precision medicine is often associated with genomic medicine, however aspects other than the genome should be considered and evidence is emerging that a patient’s microbiome plays a key role in shaping drug efficacy and should be considered as a factor that would enhance the field of precision medicine, complementing genetic information and improving patient outcomes 3.
The composition of a patient’s microbiome can be easily determined through DNA sequencing, providing a patient-specific fingerprint/barcode of the microbial communities present in a sample and an opportunity to assess whether the patient has a dysbiotic microbiome or is missing key microorganisms that would enhance the success of therapeutic treatment.
The role of the microbiome in cancer treatment is receiving significant attention and there is growing evidence that suggests the microbiome plays an important part in the way a patient responds to cancer therapies, and several studies have shown that the gut microbiome has an effect on the response to checkpoint inhibitor immunotherapies 11, 12, for example, improved efficacy of anti-PD1 treatment, with increased antitumor T cell responses, was seen in mice transplanted with faecal microbiota from responding patients12.
Although preliminary, these observations suggest that the gut microbiota might offer the opportunity to identify patients who are more likely to respond to treatment, to enhance existing therapeutics, and to develop novel therapeutic strategies.
Pharmacomicrobiomics is defined as the effect of a patient’s microbiome on drug disposition, action, and toxicity 13 and looks at the interaction between xenobiotics, or foreign compounds, and the gut microbiome.
According to the PharmacoMicrobiomics database, which aims to collect, classify and cross-reference known drug-microbiome interactions, over 60 drugs have been identified as having microbiome interactions 14. It is likely that many more interactions will be uncovered, and the plasticity of the microbiome will mean that drug interactions are dynamic 3.
Furthermore, the efficacy of prodrugs (drugs that are metabolised into a pharmacologically active version after administration) is likely to be dependent on the microbiome, which has the potential to improve or worsen the outcome for the patient. The microbiome itself can also be altered by some medications, for example antibiotics. Pathogen-specific antimicrobials such as species-specific enzyme inhibitors may have a beneficial clinical impact, and revisiting the use of bacteriophages as an alternative to antibiotics should be considered as a method of removing specific species from a community, helping to restructure the consortium 15.

Computational microbiome studies
Although it is evident that the microbiome is linked to disease studies in this field are challenging due to processing and understanding the huge amounts of data needed to find strong links between specific genes or microorganisms and disease. Microsoft has partnered with Cambridge-based Eagle Genomics to scale up its artificial intelligence platform for microbiome research, using machine learning and cloud computing to analyse large and complex datasets aiming to help users gain insights into microbiome research for healthcare.
What could the future hold?
Microbiome research looks set to continue as a high growth area, as demonstrated by an increase in the number of published articles on the topic of cancer-microbiome interactions of nearly 2000% between 2005 and 2015. There is still much research to be done to understand the functional links between the microbiome and disease, but this work is opening up a vast range of opportunities in human health. Future opportunities in microbiome research could potentially include areas such as:
- Accurate control over manipulation of microbiome, e.g. through the use of phage therapy
- The microbiome as an integral part of precision medicine, with fingerprinting of a patients microbiome becoming a complementary tool to determine how likely a patient will be to respond to treatment
- Enhancement of existing therapies by co-administration of rationally designed, patient specific, precision probiotics
- The microbiome as a drug delivery system, for example engineered probiotics containing genetic circuits to enable production of therapeutic proteins upon detection of disease biomarkers enabling on-demand drug release 10
- An expanded library of prebiotics to stimulate a diverse range of beneficial bacteria
References
- Amon P, Sanderson I. What is the microbiome? Archives of Disease in Childhood — Education and Practice. 2017;102:257 – 260
- Hooper LV, Littman DR, Macpherson AJ. Interactions Between the Microbiota and the Immune System. Science. 2012; 336(6086):1268 – 73.
- Kuntz TM, Gilbert JA. Introducing the Microbiome into Precision Medicine. Trends Pharmacol Sci. 2017; 38(1): 81- 91.
- Huttenhower C, Gevers D, Knight R et al. The Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012; 486(7402): 207 – 14.
- Lozupone CA, Stombaugh JI, Gordon, JI, Jansson, JK, Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature. 2012; 489(7415): 220 – 230.
- Sampson TR, Mazmanian SK. Control of brain development, function, and behavior by the microbiome. Cell Host Microbe. 2015; 17(5): 565 – 576.
- Bravo JA, Forsythe P, Chew MV, et al. Ingestion of Lactobacillus strain regulates emotional behavior and central GABA receptor expression in a mouse via the vagus nerve. Proc Natl Acad Sci USA. 2011; 108(38): 16050 – 16055.
- Savignac HM, Kiely B, Dinan TG, Cryan JF. Bifidobacteria exert strain-specific effects on stress-related behavior and physiology in BALB/c mice. Neurogastroenterol Motil. 2014; 26(11):1615 – 1627.
- Savignac HM, Tramullas M, Kiely B, Dinan TG, Cryan JF. Bifidobacteria Modulate Cognitive Processes in an Anxious Mouse Strain. Behav Brain Res. 2015; 287:59 – 72.
- Mimee M, Citorik RJ, Lu TK. Microbiome therapeutics — Advances and challenges. Adv Drug Deliv Rev. 2016; 105(Pt A):44 – 54.
- Chaput N, Lepage P, Coutzac C, et al. Baseline gut microbiota predicts clinical response and colitis in metastatic melanoma patients treated with ipilimumab. Ann Oncol. 2017; 28(6):1368 – 1379.
- Matson V, Fessler J, Bao R, et al. The commensal microbiome is associated with anti-PD‑1 efficacy in metastatic melanoma patients. Science. 2018; 359(6371):104 – 108.
- Rizkallah MR, Saad R, Aziz RK. The Human Microbiome Project, Personalized Medicine and the Birth of Pharmacomicrobiomics. Current Pharmacogenomics and Personalized Medicine. 2010; 8(3):182- 193.
- Rizkallah MR, Gamal-Eldin S, Saad R, Aziz RK. The PharmacoMicrobiomics Portal: A Database for Drug-Microbiome Interactions. Current Pharmacogenomics and Personalized Medicine. 2012; 10(3): 195- 303.
- Lin DM, Koskella B, Lin HC. Phage therapy: An alternative to antibiotics in the age of multi-drug resistance. World J Gastrointest Pharmacol Ther. 2017; 8(3):162 – 173.
- Microsoft Signs its First Microbiome Partnership. Available at: https://www.labiotech.eu/medic… microbiome and cancer: what’s all the fuss about? Available at: https://www.cancerresearchuk.org/funding-for-researchers/research-features/2017 – 06-13-the-microbiome-and-cancer-whats-all-the-fuss-about
Enjoyed this article? Keep reading more expert insights...
CPI ensures that great inventions gets the best opportunity to become a successfully marketed product or process. We provide industry-relevant expertise and assets, supporting proof of concept and scale up services for the development of your innovative products and processes.